Institut Pasteur Isolates Strains Of Coronavirus 2019-nCoV Detected In France
With the whole viral genome of coronavirus 2019-nCoV having recently been sequenced at the Institut Pasteur, the isolation of strains of coronavirus 2019-nCoV detected in France has now been successfully finalized, in a very short space of time, using the samples taken from the first confirmed French cases.
Coronavirus 2019-nCoV, responsible for the cases of pneumonia that emerged in China, differs from two other viruses that are well known for causing respiratory outbreaks in recent years: the SARS-CoV virus, responsible for the SARS outbreak in 2003, and MERS-CoV, responsible for an outbreak that has been under way since 2012 in the Middle East.
The Institut Pasteur was actively involved in tackling these previous outbreaks, which yielded valuable lessons for the current situation. “For both SARS-CoV and MERS-CoV, cells known as Vero E6 were identified to culture the two coronaviruses,” explains Sylvie van der Werf, Director of the National Reference Center (CNR) for Respiratory Viruses at the Institut Pasteur. “In January 2020, we brought them out of our collection, which is kept under strictly controlled conditions, so that we would be ready as soon as we detected a positive sample for coronavirus 2019-nCoV.”
Extremely rapid growth of the virus in culture
The Institut Pasteur was therefore well prepared, and on Friday January 24, 2020, the very day that the first cases were confirmed, it began the process of culturing the samples that had tested positive for the virus on Vero E6 cells.
“Using detection methods, we had observed a high viral load in the samples taken from the patients in hospital in Paris. This enabled us to identify which samples should be cultured first,” says Sylvie Behillil, Deputy Director of the CNR at the Institut Pasteur.
The viruses continued to be cultured over the weekend of January 25-26, 2020. By the morning of Monday January 27, the culture had already grown.
“We didn’t think that it would grow so quickly,” continues Sylvie Behillil. The rapid growth of the culture may be explained by “the high viral load in the samples,” but also by “the quality of the samples,” adds Vincent Enouf, Deputy Director of the CNR at the Institut Pasteur.
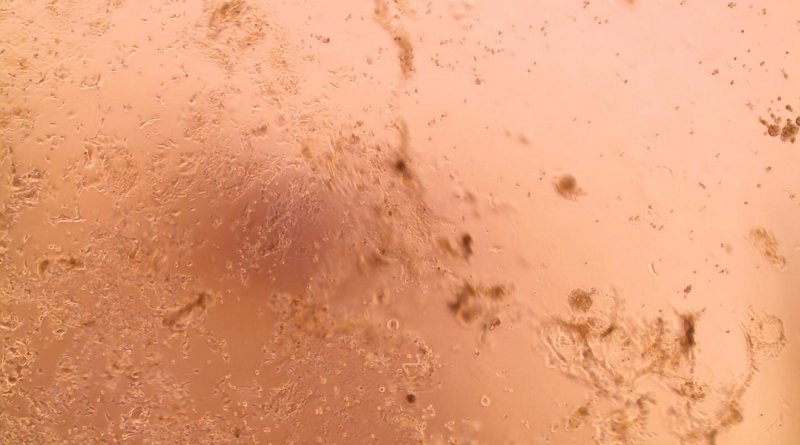
“We could see the cells becoming damaged and then grouping together, which can indicate that they have been infected. But we did not observe this cytopathic effect for all the inoculated samples; that reassured us that we had managed to isolate the strains, and this was then confirmed by additional analyses.”
Virus 2019-nCoV now available for research
Now that the Institut Pasteur’s scientists have access to coronavirus 2019-nCoV, they can set out to improve scientific knowledge about the virus.
Research will focus on four main areas.
- Serology. Analyzing antigen-antibody reactions based on the antibodies found in patients’ blood serum, and developing an effective serology test to screen for the infection among the population.
NB: This is not a rapid diagnostic test for hospital use; it is a test to identify seroconversion in the population. - Development of specific treatments. Testing known antiviral molecules that act on the replication cycle of some viruses to assess their therapeutic or even prophylactic potential, and looking for antibodies that may have therapeutic applications.
- Vaccination. Based on the virus, developing vaccine approaches that have already been explored for other viruses – Ebola, MERS-CoV and SARS-CoV -, with the aim of proposing a vaccine candidate.
- Viral pathogenesis. Understanding how the virus works, how it replicates and interacts with the cell and the host organism, to gain a clearer picture of its pathogenic nature and identify biomarkers for infection or new targets for the development of treatments.